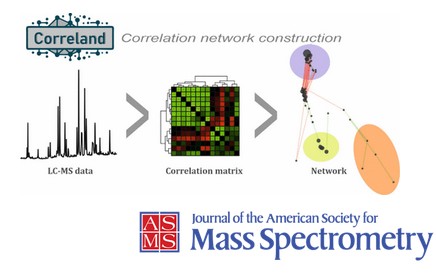
Our department presents a novel approach to LC-MS data processing based on correlation analysis. Complex spectral data are transformed into correlation matrices, enabling the identification of related signals, which are subsequently represented as intuitive network structures. This workflow provides a systematic way to move from raw, high-dimensional data to an interpretable representation that captures underlying relationships between detected features.
By focusing on correlations across multiple samples, the method allows grouping of signals that likely originate from the same compound, metabolic pathway, or chemically related species. The resulting network representation highlights clusters of co-varying features and supports the detection of patterns that would be difficult to recognize using conventional analysis strategies. This is particularly valuable for large-scale metabolomic and chemical datasets, where manual interpretation becomes impractical and important connections may remain hidden.
In addition, the approach improves robustness against noise and experimental variability by emphasizing reproducible relationships rather than individual signal intensities. The network-based view also provides a flexible platform for integrating additional layers of information, such as annotation data or structural insights, further enhancing the biological and chemical interpretability of the results.
The results of our work have been published in the Journal of the American Society for Mass Spectrometry. The study introduces a new tool for correlation network construction that can significantly contribute to deeper and more reliable data analysis in mass spectrometry and systems biology.
J. Am. Soc. Mass Spectrom. 2026.
https://doi.org/10.1021/jasms.6c00065